
GPS 103: GMM Activity Classification and Validation
Source:vignettes/gps-103-activity-and-validation.Rmd
gps-103-activity-and-validation.RmdOverview
This tutorial covers the next stage after cleaning and metric generation:
- classify each GPS row as
activeorinactivewith GMM, - smooth state flicker with HMM-based temporal smoothing,
- manually label rows in an interactive app,
- compare model predictions against labels,
- rank parameter settings by agreement.
Why This Workflow?
State classification is iterative. A clear validation loop gives better decisions:
- You can inspect how model assumptions map to real movement behaviour.
- You can use manual labels as a transparent ground-truth reference.
- You can tune parameters with a reproducible and auditable process.
- You can report uncertainty and model agreement directly.
2) Build a small reproducible example
set.seed(202)
timestamps <- seq(
from = as.POSIXct("2024-06-01 00:00:00", tz = "UTC"),
by = "10 min",
length.out = 3 * 24 * 6
)
animal_info <- tibble(
sensor_id = c("A01", "A02"),
lon0 = c(132.310, 132.316),
lat0 = c(-14.462, -14.467)
)
gps_list <- list()
for (i in seq_len(nrow(animal_info))) {
gps_list[[i]] <- tibble(
sensor_id = animal_info$sensor_id[i],
datetime = timestamps,
lon = animal_info$lon0[i] + cumsum(rnorm(length(timestamps), 0, 0.00020)),
lat = animal_info$lat0[i] + cumsum(rnorm(length(timestamps), 0, 0.00016))
)
}
gps <- bind_rows(gps_list)
gps <- grz_clean(
data = gps,
steps = c("errors", "speed_fixed", "denoise"),
max_speed_mps = 4,
verbose = FALSE
)How this works in practice:
- Start from already cleaned GPS rows with stable identifiers.
- Keep example windows small enough to iterate quickly while tuning.
Common pitfalls and checks:
- Pitfall: tuning on too broad a window early on.
Check: begin with one animal and a short period, then scale up.
3) Run GMM active/inactive classification
grz_classify_activity_gmm() fits a two-component
Gaussian mixture on movement features, then can apply optional HMM
smoothing to reduce rapid state flicker.
gps_states <- grz_classify_activity_gmm(
data = gps,
groups = "sensor_id",
feature_set = "adaptive",
adaptive_window_mins = "auto",
adaptive_window_mult = 4,
adaptive_window_min_mins = 30,
smoothing = "hmm",
hmm_self_transition = 0.98,
verbose = FALSE
)
state_view <- gps_states %>%
as_tibble() %>%
select(sensor_id, datetime, lon, lat, activity_state_gmm, inactive_prob_gmm)
state_view
#> # A tibble: 864 × 6
#> sensor_id datetime lon lat activity_state_gmm
#> <chr> <dttm> <dbl> <dbl> <chr>
#> 1 A01 2024-06-01 00:00:00 132. -14.5 NA
#> 2 A01 2024-06-01 00:10:00 132. -14.5 NA
#> 3 A01 2024-06-01 00:20:00 132. -14.5 active
#> 4 A01 2024-06-01 00:30:00 132. -14.5 active
#> 5 A01 2024-06-01 00:40:00 132. -14.5 active
#> 6 A01 2024-06-01 00:50:00 132. -14.5 active
#> 7 A01 2024-06-01 01:00:00 132. -14.5 active
#> 8 A01 2024-06-01 01:10:00 132. -14.5 active
#> 9 A01 2024-06-01 01:20:00 132. -14.5 active
#> 10 A01 2024-06-01 01:30:00 132. -14.5 inactive
#> # ℹ 854 more rows
#> # ℹ 1 more variable: inactive_prob_gmm <dbl>
state_more <- setdiff(names(gps_states), names(state_view))
cat(
"i",
length(state_more),
"more columns:",
paste(head(state_more, 10), collapse = ", "),
if (length(state_more) > 10) ", ..." else "",
"\n"
)
#> i 10 more columns: lon_raw, lat_raw, step_dt_s, step_m, speed_mps, bearing_deg, turn_rad, cum_distance_m, net_displacement_m, activity_component_gmmKey state variables in this workflow:
-
activity_state_gmm: finalactive/inactivestate after optional smoothing. -
inactive_prob_gmm: inactivity probability (smoothed ifsmoothing = "hmm"). -
activity_component_gmm: assigned mixture component id.
How this works in practice:
- GMM separates low-movement and high-movement behaviour from engineered features.
- Optional HMM smoothing adds temporal consistency to state sequences.
Common pitfalls and checks:
- Pitfall: assuming one setting is always correct across
deployments.
Check: tune and validate on representative subsets. - Pitfall: class imbalance leading to misleading accuracy.
Check: inspect class proportions by day and animal.
gps_states %>%
as_tibble() %>%
ggplot(aes(x = lon, y = lat, color = activity_state_gmm)) +
geom_point(alpha = 0.7, size = 1.3) +
facet_wrap(~sensor_id, scales = "free") +
scale_color_manual(values = c(active = "#1a9641", inactive = "#d7191c")) +
labs(
title = "GMM Activity State Classification in Space",
x = "Longitude",
y = "Latitude",
color = "State"
) +
theme_minimal()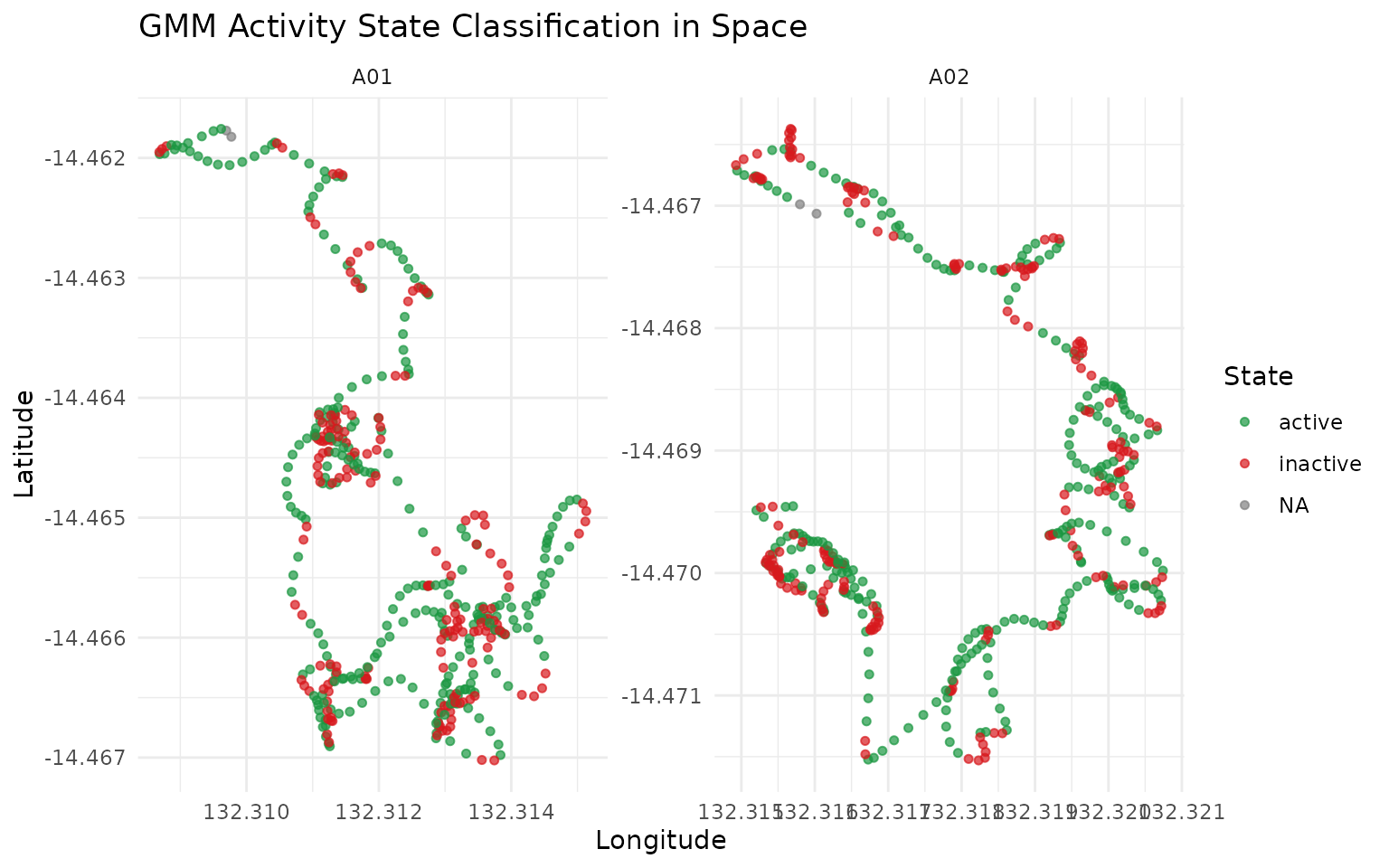
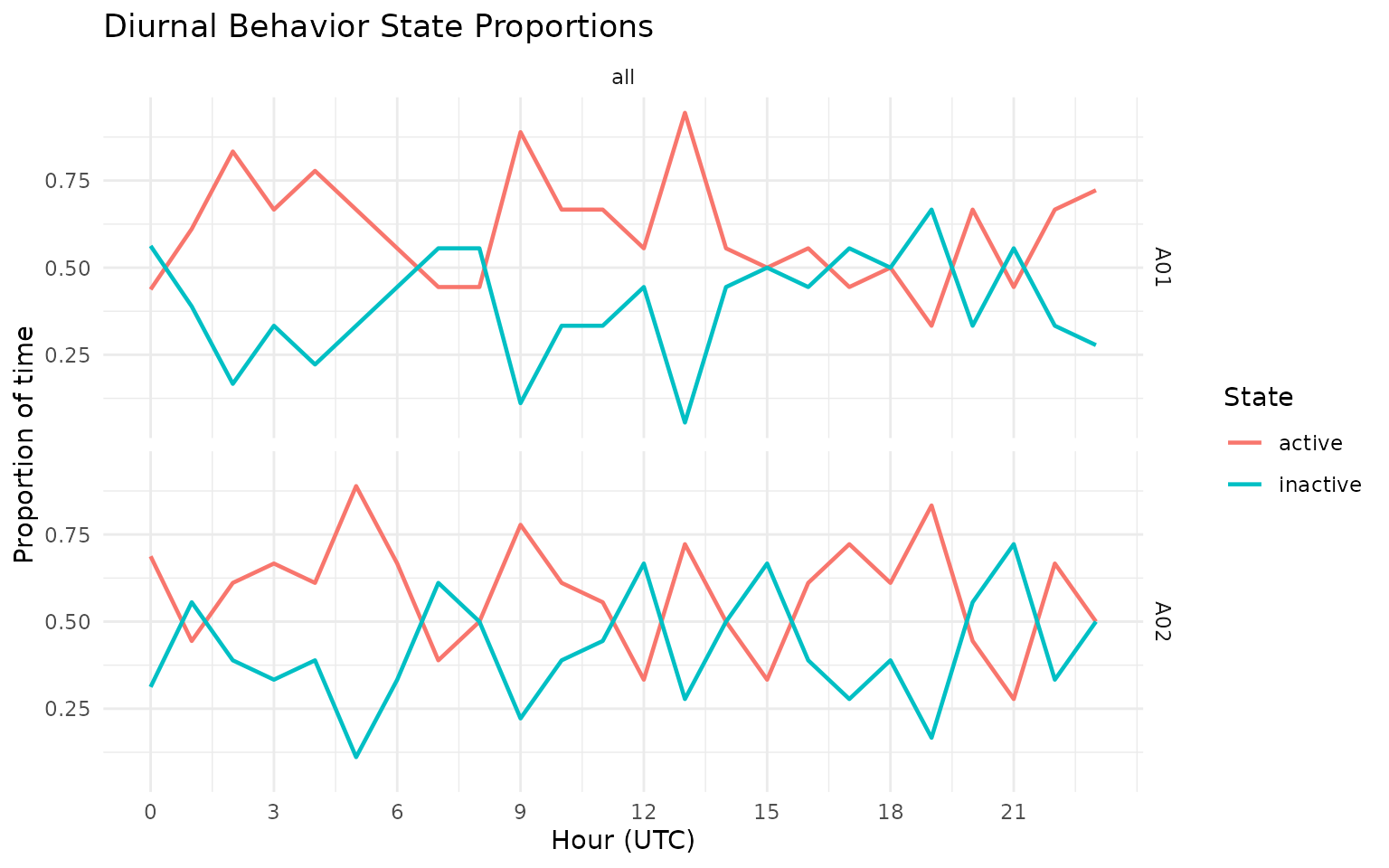
grz_plot_diurnal_states(
data = gps_states,
state_col = "activity_state_gmm",
group_col = "sensor_id",
plot_type = "line"
)
gps_states %>%
as_tibble() %>%
mutate(day = as.Date(datetime)) %>%
group_by(sensor_id, day, activity_state_gmm) %>%
summarise(n = n(), .groups = "drop") %>%
group_by(sensor_id, day) %>%
mutate(prop = n / sum(n)) %>%
ungroup() %>%
ggplot(aes(x = day, y = prop, fill = activity_state_gmm)) +
geom_col(position = "stack") +
facet_wrap(~sensor_id) +
scale_fill_manual(values = c(active = "#1a9641", inactive = "#d7191c")) +
labs(
title = "Daily Proportion of Active vs Inactive States",
x = "Day",
y = "Proportion of fixes",
fill = "State"
) +
theme_minimal()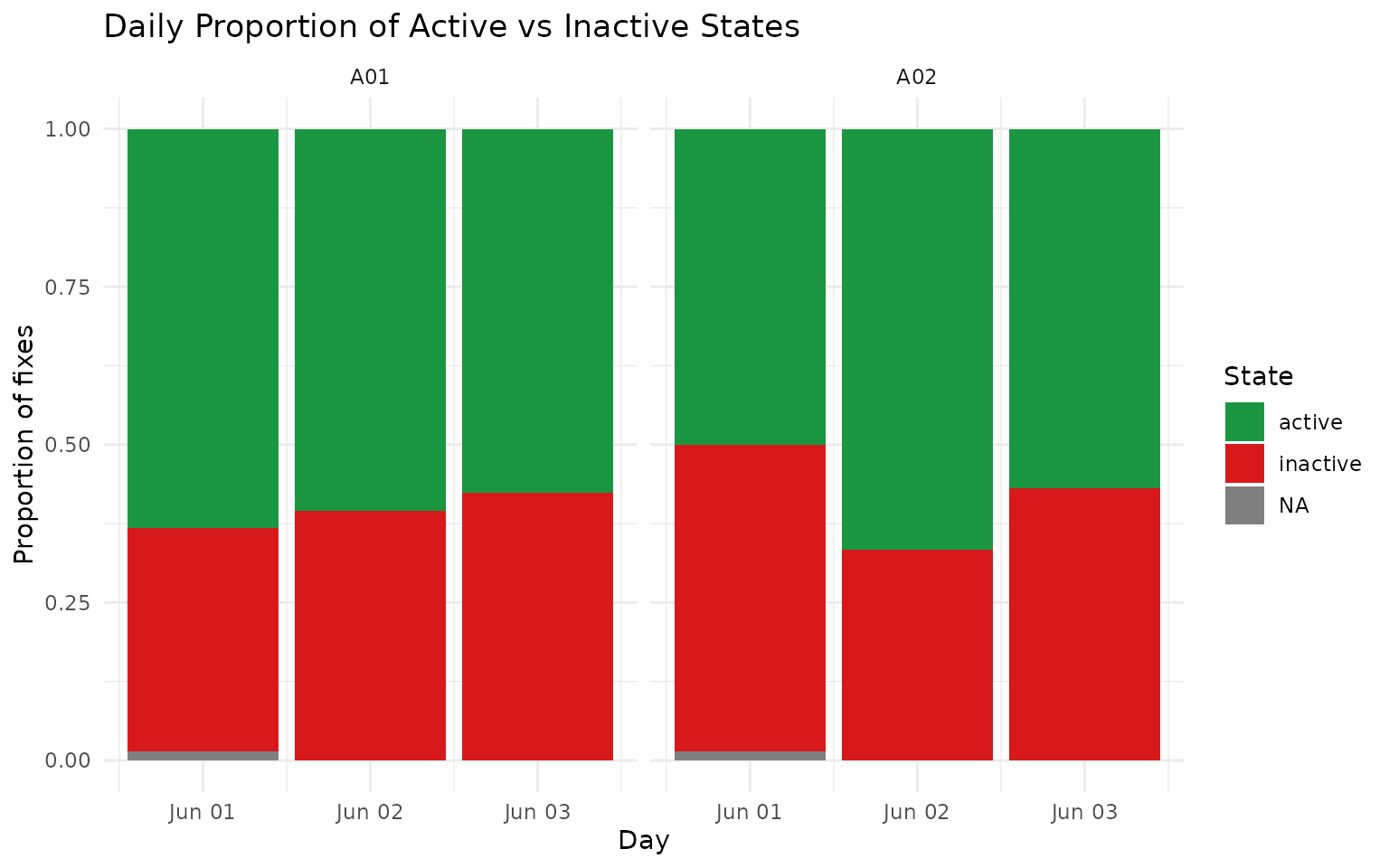
4) Add manual labels (interactive app)
Use this in an interactive R session to create or edit a
label column.
gps_states$point_id <- paste(
gps_states$sensor_id,
format(gps_states$datetime, "%Y%m%d%H%M%S", tz = "UTC"),
seq_len(nrow(gps_states)),
sep = "_"
)
labelled <- grz_label_gps_states(
data = gps_states,
lon = "lon",
lat = "lat",
time = "datetime",
id = "point_id",
color_by = "sensor_id",
initial_label_col = "label",
start_day_offset = 0L,
time_window = "week",
n_animals = 1L,
animal_col = "sensor_id"
)If you save that output to CSV, you can reload it later and continue editing:
# data.table::fwrite(labelled, "labelled_states.csv")
# labelled <- data.table::fread("labelled_states.csv")
# labelled$datetime <- as.POSIXct(labelled$datetime, tz = "UTC")How this works in practice:
- Manual labels provide ground truth for calibration and validation.
- Labels should be saved with stable IDs so they can be reused across sessions.
Common pitfalls and checks:
- Pitfall: relabelling the same points without versioning.
Check: keep dated label files or append a reviewer/version column. - Pitfall: using ambiguous behaviour windows.
Check: mark uncertain points asNAinstead of forcing a label.
5) Demonstrate a prediction-vs-label comparison
For a fully reproducible vignette, we create a synthetic
label column by copying predictions and flipping a random
subset of rows.
In your project, replace this with manual labels from
grz_label_gps_states().
labelled <- gps_states
labelled$label <- labelled$activity_state_gmm
set.seed(303)
flip_n <- floor(0.1 * nrow(labelled))
flip_idx <- sample(seq_len(nrow(labelled)), size = flip_n)
labelled$label[flip_idx] <- ifelse(
labelled$label[flip_idx] == "active",
"inactive",
"active"
)
label_view <- labelled %>%
as_tibble() %>%
select(sensor_id, datetime, activity_state_gmm, label, inactive_prob_gmm)
label_view
#> # A tibble: 864 × 5
#> sensor_id datetime activity_state_gmm label inactive_prob_gmm
#> <chr> <dttm> <chr> <chr> <dbl>
#> 1 A01 2024-06-01 00:00:00 NA NA NA
#> 2 A01 2024-06-01 00:10:00 NA NA NA
#> 3 A01 2024-06-01 00:20:00 active active 0.000999
#> 4 A01 2024-06-01 00:30:00 active active 0.000256
#> 5 A01 2024-06-01 00:40:00 active inactive 0.000000155
#> 6 A01 2024-06-01 00:50:00 active active 0.000000120
#> 7 A01 2024-06-01 01:00:00 active active 0.000000132
#> 8 A01 2024-06-01 01:10:00 active active 0.000000322
#> 9 A01 2024-06-01 01:20:00 active active 0.000708
#> 10 A01 2024-06-01 01:30:00 inactive inactive 1.000
#> # ℹ 854 more rows
label_more <- setdiff(names(labelled), names(label_view))
cat(
"i",
length(label_more),
"more columns:",
paste(head(label_more, 10), collapse = ", "),
if (length(label_more) > 10) ", ..." else "",
"\n"
)
#> i 12 more columns: lon, lat, lon_raw, lat_raw, step_dt_s, step_m, speed_mps, bearing_deg, turn_rad, cum_distance_m , ...How this works in practice:
- Truth labels and predicted states are compared on the same rows.
- Only valid binary states (
active,inactive) are included for scoring.
6) Tune parameters and rank the top 3 settings
grid <- tidyr::expand_grid(
adaptive_window_mult = c(3, 4, 5),
hmm_self_transition = c(0.95, 0.975, 0.99)
)
results <- tibble()
for (i in seq_len(nrow(grid))) {
pred <- grz_classify_activity_gmm(
data = labelled,
groups = "sensor_id",
feature_set = "adaptive",
adaptive_window_mins = "auto",
adaptive_window_mult = grid$adaptive_window_mult[[i]],
adaptive_window_min_mins = 30,
smoothing = "hmm",
hmm_self_transition = grid$hmm_self_transition[[i]],
verbose = FALSE
)
comparison <- tibble(
truth = tolower(trimws(as.character(labelled$label))),
pred = tolower(trimws(as.character(pred$activity_state_gmm)))
) %>%
filter(truth %in% c("active", "inactive")) %>%
filter(pred %in% c("active", "inactive"))
n_compared <- nrow(comparison)
accuracy <- if (n_compared == 0L) {
NA_real_
} else {
mean(comparison$truth == comparison$pred)
}
results <- bind_rows(
results,
tibble(
adaptive_window_mult = grid$adaptive_window_mult[[i]],
hmm_self_transition = grid$hmm_self_transition[[i]],
n_compared = n_compared,
accuracy = accuracy
)
)
}
top3 <- results %>%
arrange(desc(accuracy), desc(n_compared)) %>%
slice_head(n = 3) %>%
mutate(accuracy_percent = round(100 * accuracy, 2)) %>%
select(adaptive_window_mult, hmm_self_transition, n_compared, accuracy_percent)
top3
#> # A tibble: 3 × 4
#> adaptive_window_mult hmm_self_transition n_compared accuracy_percent
#> <dbl> <dbl> <int> <dbl>
#> 1 4 0.975 860 90.1
#> 2 4 0.99 860 89.5
#> 3 4 0.95 860 88.7How this works in practice:
- A parameter grid is evaluated with a consistent GMM+HMM configuration.
- Each setting is ranked by observed agreement against manual labels.
- Top settings are candidates for holdout validation, not automatic final choices.
results %>%
mutate(accuracy_percent = 100 * accuracy) %>%
ggplot(aes(x = factor(adaptive_window_mult), y = factor(hmm_self_transition), fill = accuracy_percent)) +
geom_tile(color = "white") +
labs(
title = "GMM Accuracy Across Parameter Grid",
x = "adaptive_window_mult",
y = "hmm_self_transition",
fill = "Accuracy (%)"
) +
theme_minimal()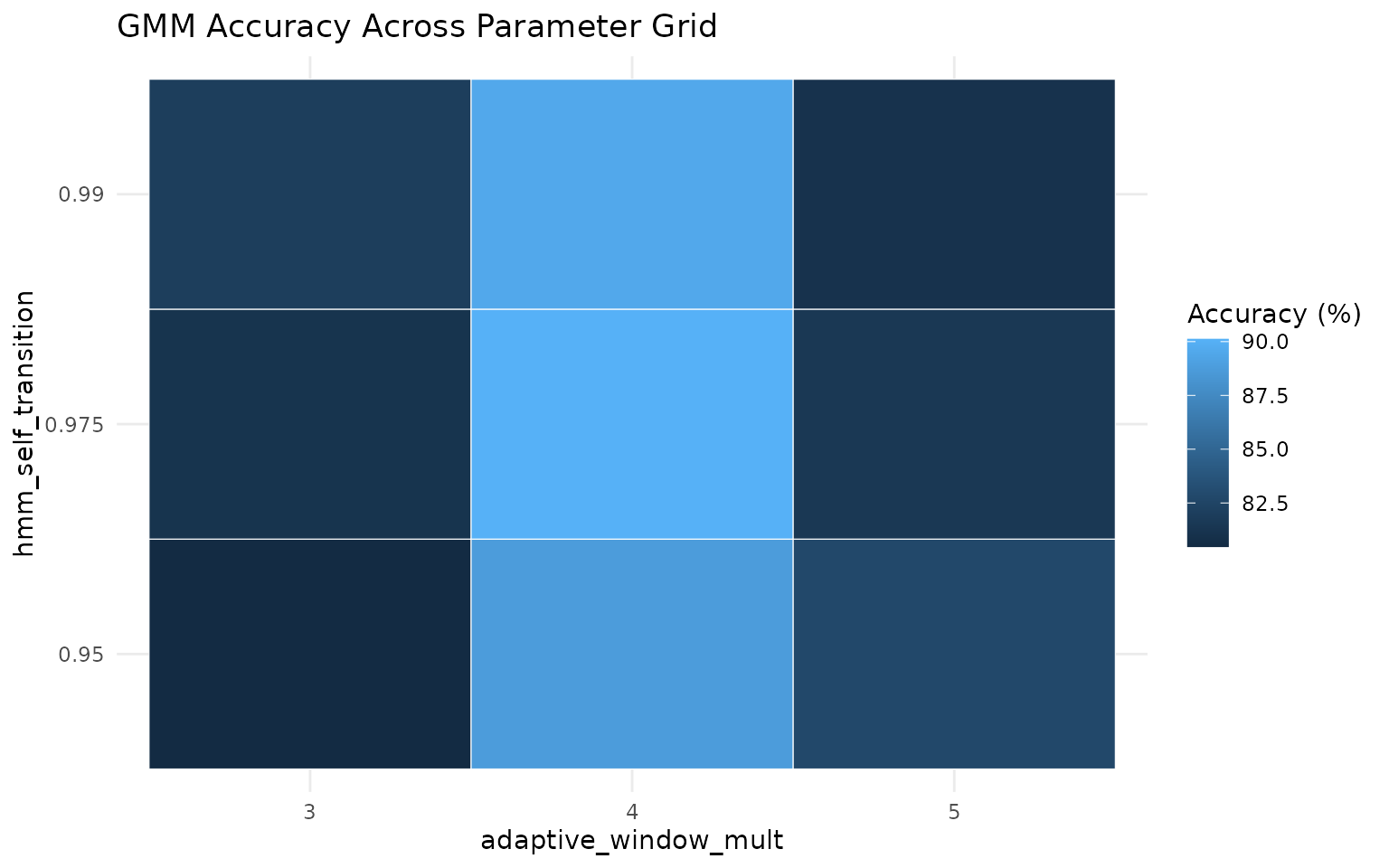
best <- top3 %>% slice(1)
best_pred <- grz_classify_activity_gmm(
data = labelled,
groups = "sensor_id",
feature_set = "adaptive",
adaptive_window_mins = "auto",
adaptive_window_mult = best$adaptive_window_mult[[1]],
adaptive_window_min_mins = 30,
smoothing = "hmm",
hmm_self_transition = best$hmm_self_transition[[1]],
verbose = FALSE
)
confusion_df <- tibble(
truth = tolower(trimws(as.character(labelled$label))),
pred = tolower(trimws(as.character(best_pred$activity_state_gmm)))
) %>%
filter(truth %in% c("active", "inactive")) %>%
filter(pred %in% c("active", "inactive")) %>%
count(truth, pred, .drop = FALSE)
confusion_df %>%
ggplot(aes(x = truth, y = pred, fill = n)) +
geom_tile(color = "white") +
geom_text(aes(label = n), size = 5) +
labs(
title = "Confusion Matrix for Best Parameter Set",
x = "Manual label",
y = "Predicted state",
fill = "Count"
) +
theme_minimal()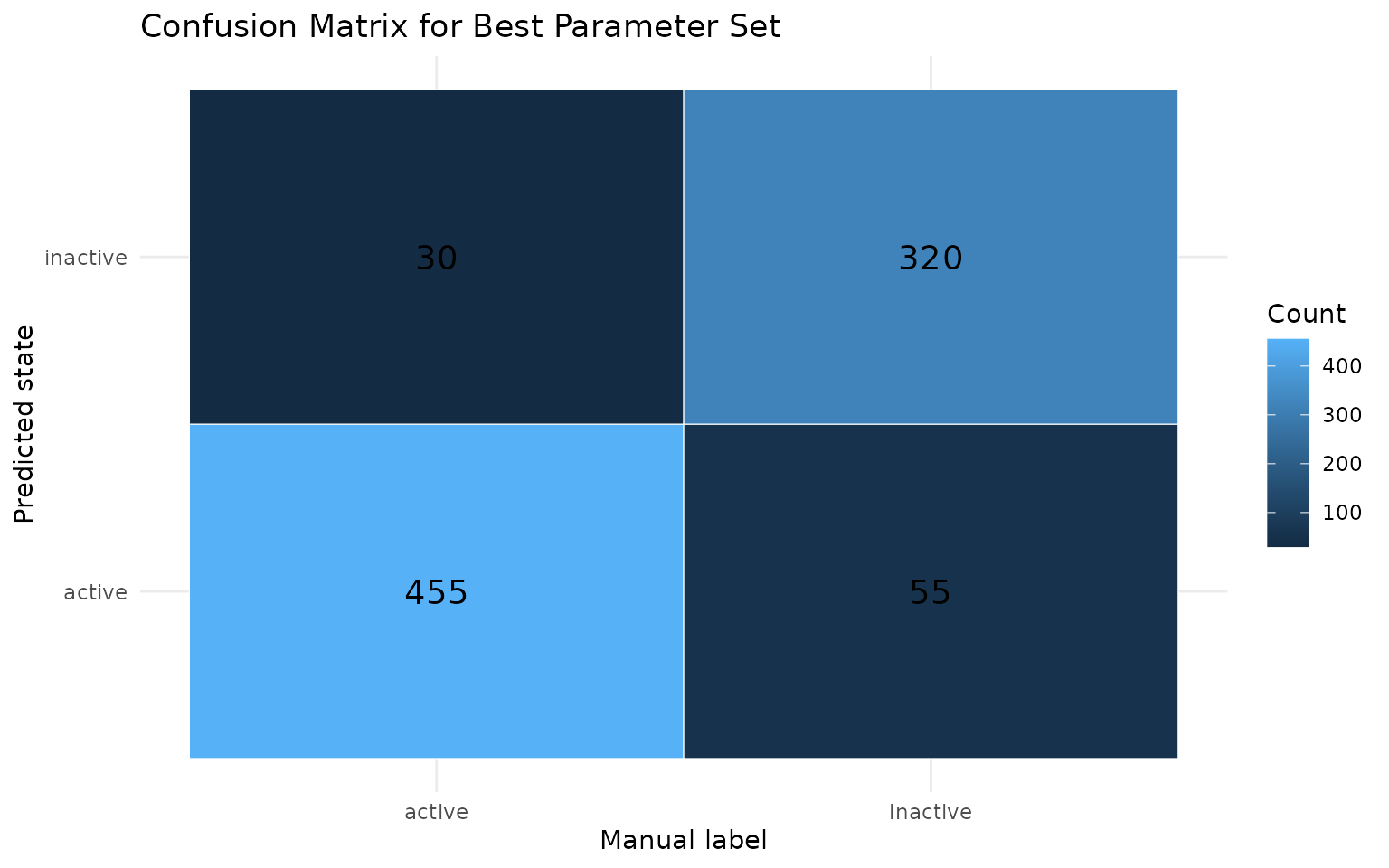
7) Optional advanced diagnostics
These helpers are useful once you have enough labelled data:
# Diurnal feature patterns for threshold setting
grz_plot_diurnal_metrics(
data = labelled,
metrics = c("step_m", "turn_rad"),
group_col = "sensor_id"
)
# Behaviour diagnostics (feature summary, transitions, bouts, PCA)
diagnostics <- grz_validate_behavior(
data = labelled,
state_col = "activity_state_gmm",
groups = "sensor_id",
pca = TRUE
)grz_validate_behavior() returns state counts,
transitions, bout summaries, and an optional PCA diagnostic for the
selected state column.